Sample specification and submission guidelines
Sample specification and submission guidelines
DNA Quality:
The DNA must be double stranded, without degradation and RNA contamination. DNA extraction by column purification method is strongly recommended (e.g. from Qiagen or any other supplier). Our service recommends avoiding phenol-chlroform in DNA extractions. The DNA can be resuspended in water or 10mM Tris-HCL, pH 7.5-8.5 (Eliminate the presence of EDTA in your samples).
The indicator of sample purity, OD260/280=1.8-2.0 and OD260/230≥1.8, are good indicators and when possible, our service recommends that DNA should be assessed by running a gel electroforesis.
The correct quantification of gDNA is essential in library construction. Our service recommends quantifying the starting genomic material by using a fluorescence based quantification method such a Qubit or picogreen assay. These measurement tend to be specific for double stranded DNA and are preferable to nanodrop or other UV based measurement. If your lab does not have a way of making a fluorescent based measurement, please measure with whatever method is available and indicate this information when submitting your sample.
RNA Quality:
For most RNA extraction, we recommend the RNeasy kit from Qiagen, however, you may use your own lab protocols or equivalent kits if they have yield high quality, without RNA
degradation. Use of degraded RNA can result in low yield, over-representation of the 5 ́ends or failure of the protocol. RNA that has DNA contamination will result in an underestimation of the amount of RNA used, for that reason, all RNA extraction must also be accompanied by DNase treatment. Qiagen protocols allow this step to be preformed on the column. Preventing RNase contamination is mandatory requirement when handling RNA samples. The purified RNA should be eluted or dissolved in RNAse-free water.
We recommend the use of Agilent Tecnologies 2100 Bioanalyzer to check the total RNA integrity. The specification for RNA Integrity Number (RIN) value of greater than 7 is requiered. Alternatively, if Bioanalyzer is not available, please run the RNA samples on a denaturing agarose gel to confirm the integrity of rRNA peaks.
The correct quantification of RNA is essential in library construction. Our service recommend quantifying the starting material by using a fluorescence based quantification method such a Qubit or Ribogreen assay. These measurement tend to be specific for RNA and are preferable to nanodrop or other UV based measurement. If your lab does not have a way of making a fluorescent based measurement, please measure with whatever available method and indicate this information when submitting your sample.
*Formalin-fixed, paraffin-embedded (FFPE) samples, that usually are strongly degraded, it is recommended to setup a pilot to confirm that materials are suitable.
*All samples undergo quality check (QC) before going into library preparation. Make sure to include enough volume to allow for 5 uL to be used in QC. The inputs listed below should remain after the completion of this QC. The following are minimum requirements for library construction. If your sample does not comply with this guidelines, please contact us and we can discuss your specific sample requirements.
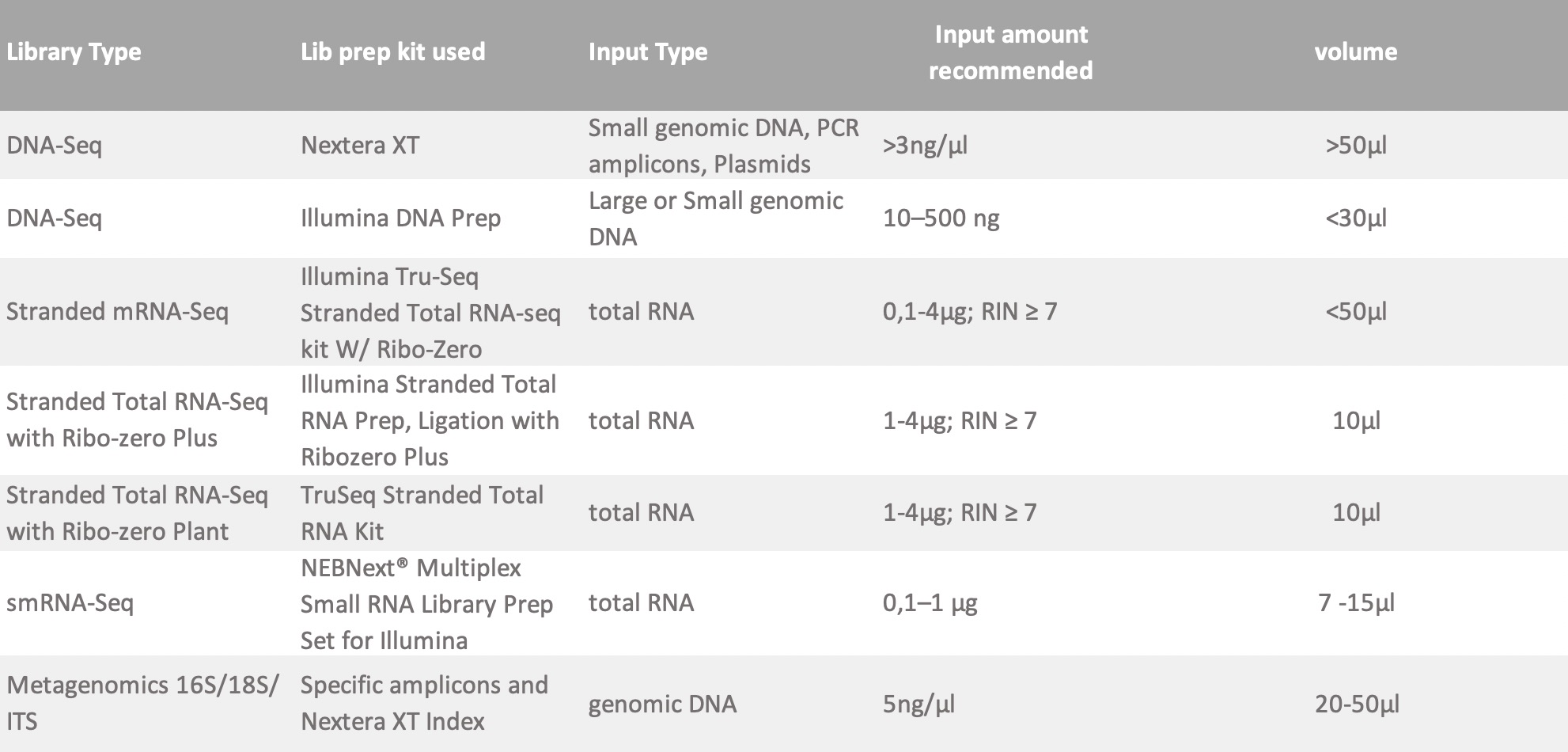
FOR PREPARED LIBRARY:
We do accept prepared libraries for sequencing. However, we do not guarantee the sequencing results for any customer prepared libraries.
Prepared libraries should be sent at ≥10 nM. A volume of 15ul at the minimum is required. Concentration should ideally be determined using fluorometric means (picogreen/Qubit). If you are pooling your libraries, the final pool should also be at at ≥10 nM with a volume of at least 15ul.
Submission Condition
TUBES:The samples should be placed in 1.5 ml microcentrifuge tubes sealed with parafilm, labelled and placed inside 50mL tubes. Make sure to write the sample number listed on your order sample sheet on the top the tube. To prevent sample tubes from moving during shipment, pack any remainning space with paper towel/tissue paper prior to sealing.
PLATES: Should you submit more than one sample, please use polypropylene PCR plates. Plates should be labeled with the order number and sample number range clearly marked, and should be sealed firmly with a foil plate seal capable of withstanding -80 C temperatures. Samples should be loaded on the plate in rows and not in columns.
*Transport should be in dry ice and send by Express company.
*Include a printed copy of the ordersheet with a description of the DNA/RNA extraction and/or enrichment protocol used. The description should incluye: Description of the sample: name, code, A260/280, concentration (ng/μl), specify quantitation methods (Pico/Ribogreen, nanodrop or/and spectrophotometre) a electrophoresis image of the EtBr-stained gel with the samples. Please include marker as a reference and clearly identify the contents and amount (in nanograms) of DNA of each lane. A Bioanalyzer profile can be incluye if it is available, mainly in RNA samples.
*Include a printed copy of your project quote in the package.
Sample Shipping to:
Laboratorio de Genómica y Ultrasecuenciación Universidad de Málaga
Edificio de Bioinnovación, 1a planta. puerta 104
Parque Tecnológico de Andalucía (PTA) C/Severo Ochoa, 34
29590 Campanillas-Málaga Spain.
Tlf. +34 951 952 780 (Attn. Josefa Gómez Maldonado)
Checklist for sample submission

All the parcel information must also be send to the contact service This email address is being protected from spambots. You need JavaScript enabled to view it.
Download guides:
